For this purpose, we re-implemented and adapted the visualization approach of our stand-alone tool GENeVis (Westenberg et al., 2007), which allows embedded visualization of all time points in the nodes. However, to detect temporal patterns in gene expression, it is necessary to consider all time points, which requires simultaneous visualization of multiple expression values. Our plugin also provides a method for visualization of time series data, because Cytoscape offers only basic support by coloring nodes (genes) based on a single time point. SpotXplore provides two methods to detect such hotspots, and allows the user to visualize the hotspots in the context of the whole network to explore regulatory interactions between differentially expressed gene sets. In this article, we present SpotXplore, a plugin for analysis and visual exploration of differentially expressed subnetworks (hotspots). The generic biological network visualization tool Cytoscape (Shannon et al., 2003) provides an excellent environment to implement combined analysis and visualization methods.  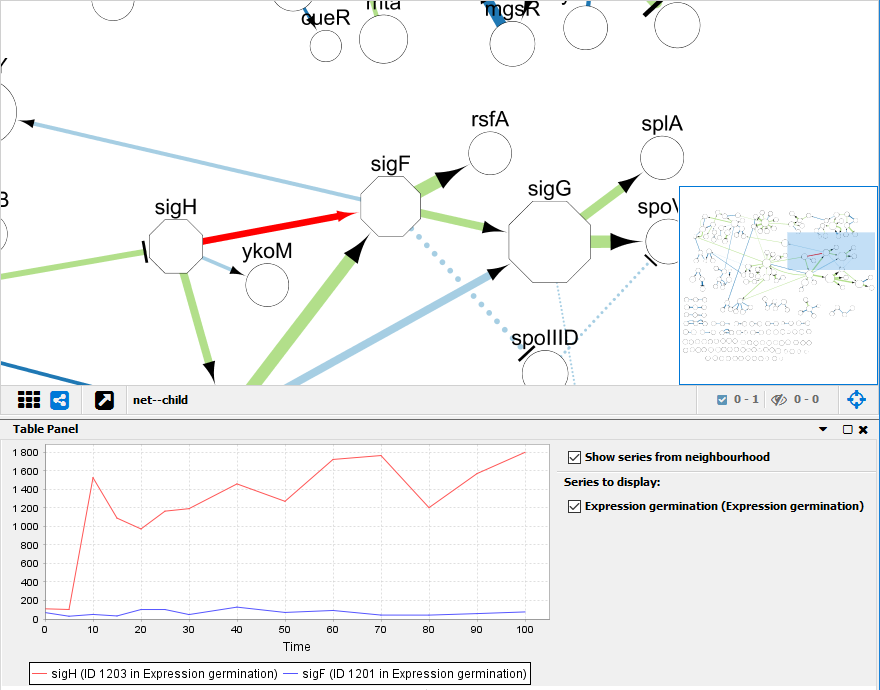
Only then, a researcher does not have to switch between various applications to obtain a comprehensive understanding of the biological processes affected in the experiments. Nevertheless, there is a need for tools that incorporate dynamics of gene expression in the networks by offering both data analysis and visualization. Many tools for visualizing the various types of biological networks have been developed (Suderman and Hallett, 2007). Insight can be obtained by analyzing the data in the context of molecular interaction networks, such as gene regulatory networks or metabolic pathways, which requires visualization in addition to statistical analysis tools. Interpretation of gene expression from time series experiments or multiple perturbation studies is a challenging task. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
